Abstract:
Background: Many molecules of interest are flexible and undergo significant shape deformation as part of their function, but most existing methods of molecular shape comparison (MSC) treat them as rigid bodies, which may lead to incorrect measure of the shape similarity of flexible molecules.
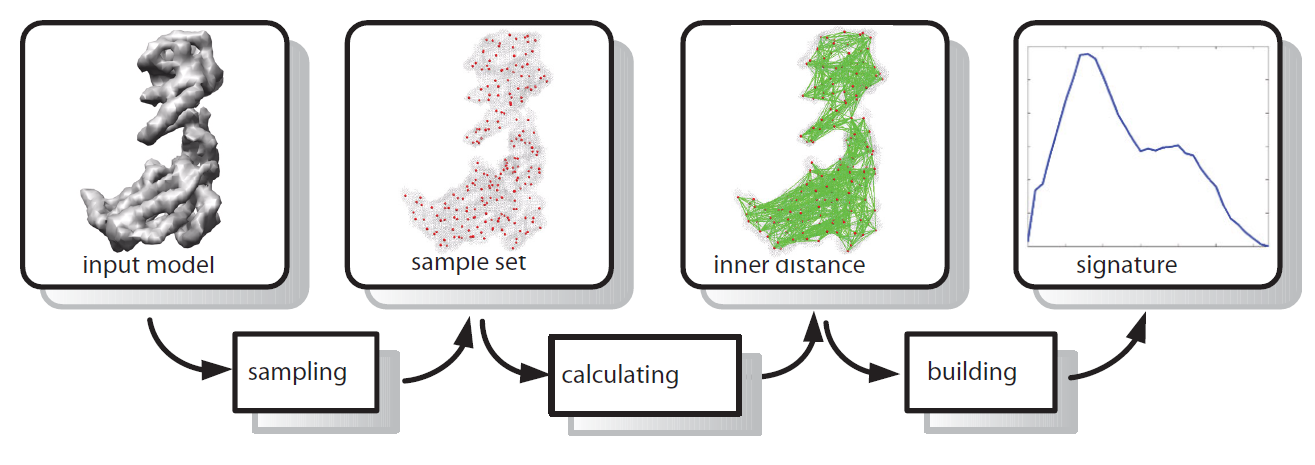
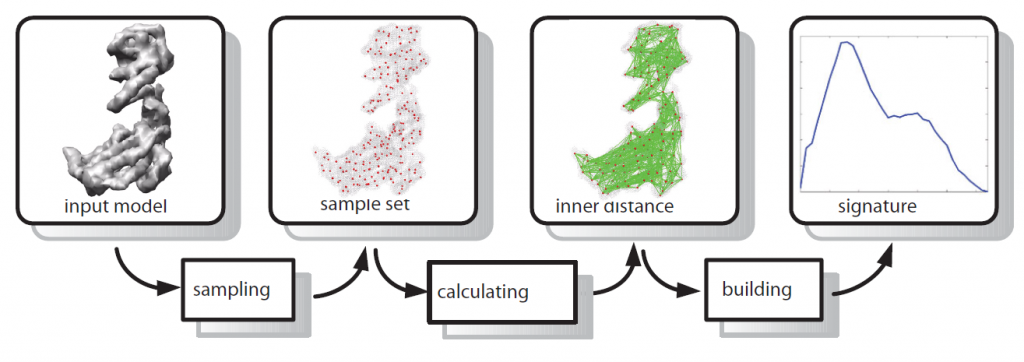
Results: To address the issue we introduce a new shape descriptor, called Inner Distance Shape Signature (IDSS), for describing the 3D shapes of flexible molecules. The inner distance is defined as the length of the shortest path between landmark points within the molecular shape, and it reflects well the molecular structure and deformation without explicit decomposition. Our IDSS is stored as a histogram which is a probability distribution of inner distances between all sample point pairs on the molecular surface. We show that IDSS is insensitive to shape deformation of flexible molecules and more effective at capturing molecular structures than traditional shape descriptors. Our approach reduces the 3D shape comparison problem of flexible molecules to the comparison of IDSS histograms.
Conclusion: The proposed algorithm is robust and does not require any prior knowledge of the flexible regions. We demonstrate the effectiveness of IDSS within a molecular search engine application for a benchmark containing abundant conformational changes of molecules. Such comparisons in several thousands per second can be carried out. The presented IDSS method can be considered as an alternative and complementary tool for the existing methods for rigid MSC. The binary executable program for Windows platform and database are available from https://engineering.purdue.edu/PRECISE/IDSS.
Link to paper:  http://www.biomedcentral.com/content/pdf/1471-2105-10-157.pdf
Download paper here
Citation:
Liu YS, Fang Y, Ramani K. IDSS: deformation invariant signatures for molecular shape comparison. BMC Bioinformatics. 2009 May 22;10:157. PMID: 19463181.