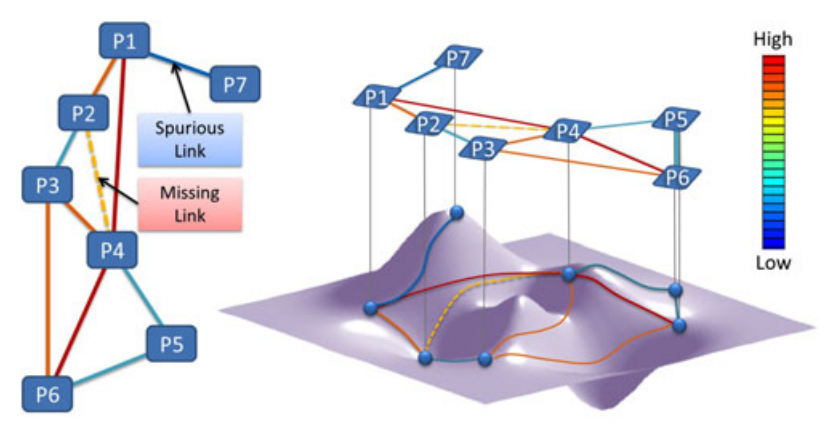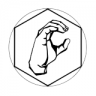
Recent developments in high-throughput technologies for measuring protein-protein interaction (PPI) have profoundly advanced our ability to systematically infer protein function and regulation. However, inherently high false positive and false negative rates in measurement have posed great challenges in computational approaches for the prediction of PPI. A good PPI predictor should be 1) resistant to high rate of missing and spurious PPIs, and 2) robust against incompleteness of observed PPI networks. To predict PPI in a network, we developed an intrinsic geometry structure (IGS) for network, which exploits the intrinsic and hidden relationship among proteins in network through a heat diffusion process. In this process, all explicit PPIs participate simultaneously to glue local infinitesimal and noisy experimental interaction data to generate a global macroscopic descriptions about relationships among proteins. The revealed implicit relationship can be interpreted as the probability of two proteins interacting with each other. The revealed relationship is intrinsic and robust against individual, local and explicit protein interactions in the original network. We apply our approach to publicly available PPI network data for the evaluation of the performance of PPI prediction. Experimental results indicate that, under different levels of the missing and spurious PPIs, IGS is able to robustly exploit the intrinsic and hidden relationship for PPI prediction with a higher sensitivity and specificity compared to that of recently proposed methods.