Our People

Principal Investigator
David Umulis, Ph.D.
Dane A. Miller Head: Weldon School of Biomedical Engineering
Professor of Biomedical Engineering
Professor of Agricultural and Biological Engineering
Email: dumulis@purdue.edu
Phone: +1 765 494-1223
Fax: +1 765 496-1115
Purdue University
Weldon School of Biomedical Engineering
206 S. Martin Jischke Drive
West Lafayette, IN 47907-2032
Staff
Xiaoguang Zhu; Ph.D.

Research Scientist
Email: zhu410@purdue.edu
Research: Determine the functional significance of feedback networks in the regulation of BMP-mediated patterning by systematically evaluating gradient dynamics, robustness, and scaling of BMP signaling in wild-type and mutant zebrafish embryos, such as bambia, sizzled, and admp.]
Publications: https://www.researchgate.net/profile/Xiaoguang-Zhu-4
Linlin Li;Ph.D.

Post Doctoral Research Assistant
Email: li2212@purdue.edu

Research: Embryonic development is a complicated phenomenon that combines cellular signaling, gene regulation, and cellular movement to form structures and tissues that presage adult physiology. Morphogens mediate the cell fate decision by its time-varying spatial distribution. In zebrafish, patterns of gene expression along the dorsal-ventral (DV) body axis are regulated by Bone Morphogenetic Proteins (BMPs). In our work, we are interested in how the cellular movements impact the formation of gradients by contributing an advective term whereby the morphogens are swept with the moving cells as they move vegetally. We developed an integrated, moving-mesh Finite Element Model whereby the cell velocities directly inform the mesh movement, providing a realistic domain for computing pattern information. Our goal is to develop a complete advection-diffusion-reaction model that incorporates all stages of zebrafish embryonic development data to investigate the mechanism in underlining BMP driven DV patterning during epiboly.
Huma Shoaib; Ph.D.

Engineering Education Researcher
Profile Link: https://www.linkedin.com/in/humashoaib/
Yixuan Liu; B.S. ABE
Lab Assistant at Purdue University
Linkin Profile: https://www.linkedin.com/in/yixuan-liu-5b0092173/
Email: liu2586@purdue.edu
Responsibilities: Support and conduct experiment with Xiaoguang zhu, Linlin Li and Matthew Thompson; zebrafish house Maintenance; and provide lab support, including imaging with confocal, preparation of reagents, lab supply ordering and participate in group meetings and journal club.
Graduate Students
Matthew Thompson; Ph.D candidate

Email: thompsmj@purdue.edu
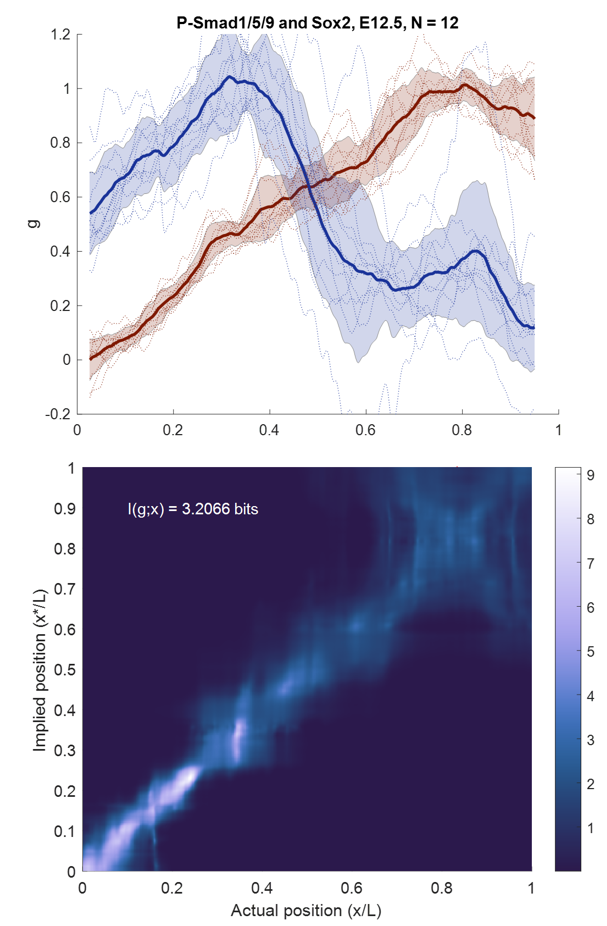
Research: The organ of Corti in the inner ear is a striking example of precise developmental patterning, where the radial pattern is faithfully repeated over 3000 times from base to apex. Each individual cell along the radial axis has a unique role, and disruption of this pattern can cause defects in hearing. I'm interested in learning how nascent cells along this axis interpret the positional information contained in transcription factor profiles before cell fate specification begins. Combining semiquantitative imaging data with tools from information theory and systems biology, I'm working on ways to decipher the complex regulatory system guiding this developmental process with an emphasis on the Bmp pathway.
Agnes Doszpoly; Ph.D student

Email: adoszpol@purdue.edu
Research Interest: Developing BTDR Fellow (Bioengineering Training in Diabetes Research), computational modeling of Type 1 Diabetes pathogenesis in zebrafish.
Nissa Jo Larson; Ph.D student

Email: larso124@purdue.edu
Research Interest: Modeling zebrafish embryo Bone Morphogenetic Protein (BMP) gradient formation, considering robustness contribution of membrane-bound BMP receptor complexes
LeRayah Neely-Brown; M.S student

Email: lneelybr@purdue.edu
Research Interest: LeRayah Neely-Brown is a first year Biomedical Engineering Master’s student. She received her BS in Mathematics and Computer Science from the University of Illinois at Chicago in 2021. Her interests lie in utilizing computational neuroscience to understand neuronal signal processing from sub-cellular to network/system-level dynamics. She aims to analyze and phenotype action potentials via biophysical and computation modeling for the vagus nerve. In her spare time, LeRayah enjoys listening to music, going to museums, and styling others’ makeup.
Alumni
Nimisha Bajaj; M.D., Ph.D.

Coadvised with Sherry Harbin and Ann Rundell

Current Position: Pediatric Resident at Nationwide Children's Hospital;
Research: Investigate the dynamics of the extracellular matrix molecules that promote and inhibit neovascularization through the mechanotransduction pathway through computational modeling and validation in vitro using a three-dimensional tissue culture system. Because the available data is largely qualitative, we are using optimal scaling and multi-objective optimization in order to fit the model parameters as well as the network structure of the experimental data.
Yan Huang; Ph.D.

Current Position: SDET at Amazon
Linkin Profile: https://www.linkedin.com/in/yanhuang1/
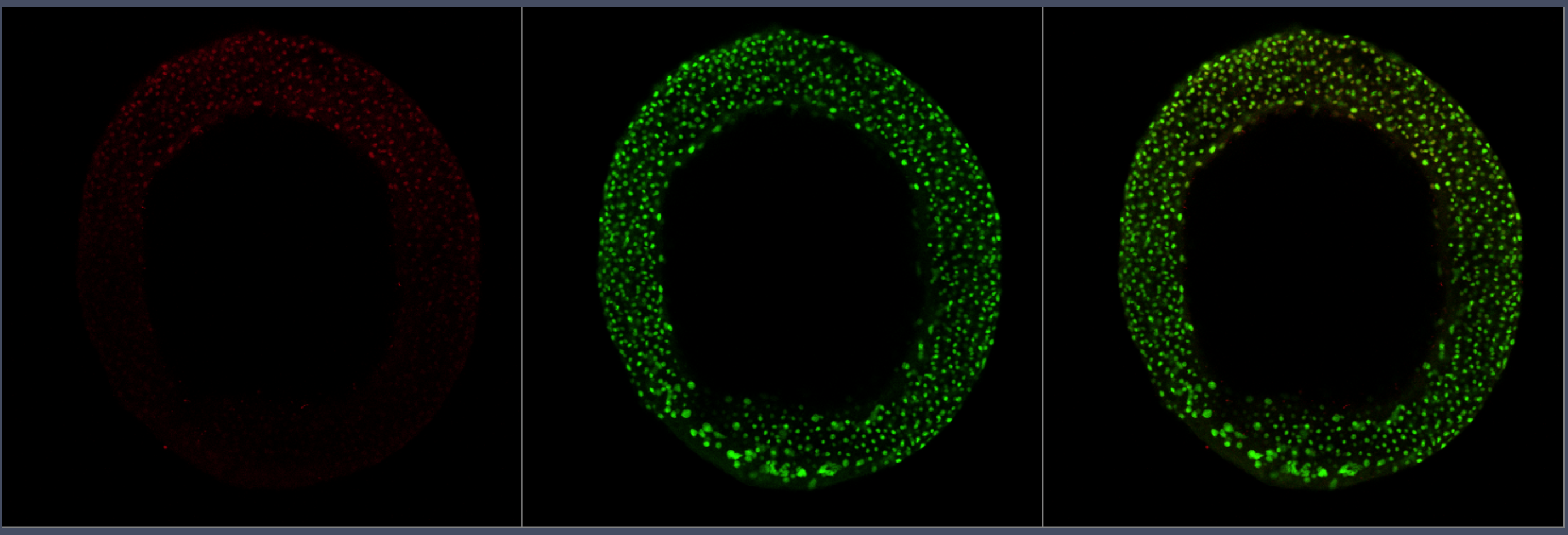
Research: Mechanism of scale invariance in Dorsal-Ventral patterning using quantitative imaging and mathematical modeling in zebrafish embryos.
Thembi Mdluli; Ph.D.

Coadvised with Ann Rundell

Current Position: Computational Biologist at The Henry M. Jackson Fundation for the Advancement of MIlitary Medicine
Linkin Profile: https://www.linkedin.com/in/thembi-mdluli-53829a1a/
Research: Model parameter optimization using disparate datasets is difficult because some datasets are more important or informative that others, and the tradeoff between these datasets are not clear. With multiple plausible models, models might have to sacrifice satisfaction of one dataset in order to improve the fitness of another. Multi-objective optimization searches for a set of solutions with a tradeoff between competing objectives which allows for a fair comparison between different models.
Joyatee Sarker; M.D., Ph.D.

Coadvised with Ann Rundell and Tamara Kinzer-Ursem
Current Position: Psychiatry Resident at Indiana University School of Medicine
Linkin Profile: https://www.linkedin.com/in/joyatee/
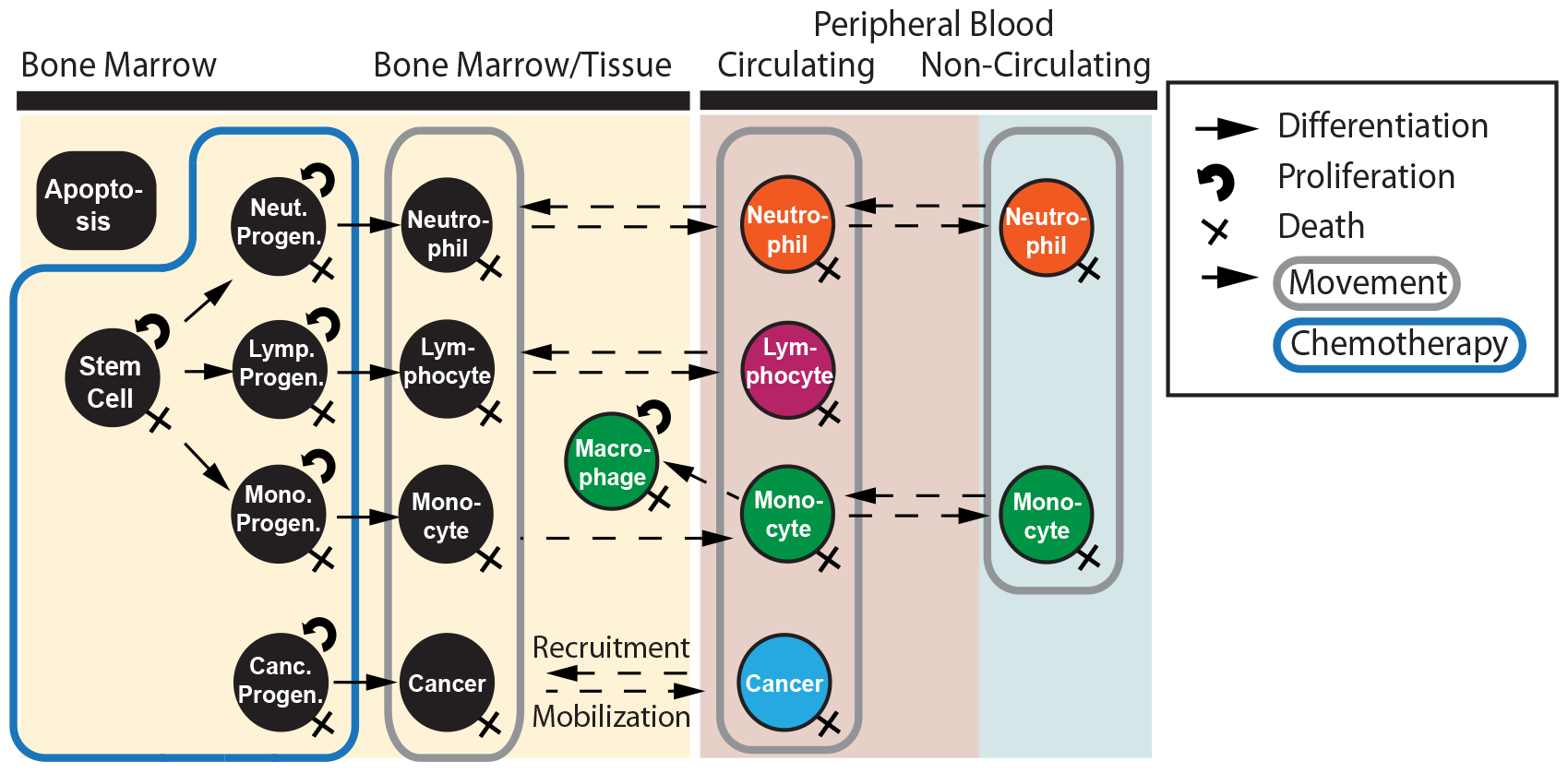
Research: Hematopoiesis within individuals differ in several ways, and Acut Myelogenous Leukemia (AML) behave differently within each patient. We have created a multi-linage computational model of the hematopoietic system, and simulated 22,796 virtual patients using unique parameter sets. Our goal is to predict the treatment schedule for individual patients' optimal recovery from leukemia.
Md. Shahriar Karim; M.S. ABE, M.S. ECE, Ph.D.

Current Position: Assistant Professor of North South University
Profile Link: http://ece.northsouth.edu/people/md-shahriar-karim/
Research Interest: Quantitative System Biology, Modeing of Dynamical Systems, Estimation and Detection Theory and Computational Biology (Cited from: http://ece.northsouth.edu/people/md-shahriar-karim/)
Michael Pargett; Ph.D.
Current Position: Project Scientist
Profile Link: https://www.mcb.ucdavis.edu/faculty-labs/albeck/aboutlab.htm
James Hengenius; Ph.D.

Current Position: Postdoc at University of Pittsburgh
Profile Link: http://huppertlab.net/people/
Joseph Zinski; Ph.D.

Co Advised with MC Mullins at Upenn
Current Position: Post Doc at Upenn
Linkin Profile: https://www.linkedin.com/in/joseph-zinski-623812a5/
Tzu-Ching Wu; Ph.D.

Current Position: PhD Candidate at Purdue Universtiy
Linkin Profile: https://www.linkedin.com/in/tzuching/
Research Website: http://tzuching1.weebly.com/
Research Fileds: Medical Image Processing, Developmental Biology, Deep Learning, Computational Biology, Multi-objective optimization (Cited from: http://tzuching1.weebly.com/)
Michelle Ingle; M.S. ABE

Linkin Profile: https://www.linkedin.com/in/michelle-ingle-9262a3152/
Juan Carvajal; M.S. ABE

Linkin Profile: https://www.linkedin.com/in/jcarvajal91/
Lukasz Burzawa; M.S. BME
Current Position: Machine Learning Engineer at Micron Technology
Linkin Profile: https://www.linkedin.com/in/lburzawa/
Yan Luo; M.S. ABE
Current Position: Software Engineer at Facebook
Linkin Profile: https://www.linkedin.com/in/luo-yan/
Wei Dou; Ph.D.

Current Position: Quantitative Researcher
Linkin Profile: https://www.linkedin.com/in/wei-dou-ph-d-37883b46/
Xu Wang; Post Doc

Current Position: Assistant Research Professor of Bioland Laboratory (Guangzhou Regenerative Medicine and Health Guangdong Laboratory)
Ye Bu; Post Doc

Current Position: Development Research Director (CTO) of Molecular Medicine Research Institute of Peking University
Bobby Madamanachi; Post Doc

Current Position: Lecturer: University of Michigan
Linkin Profile: https://www.linkedin.com/in/bobby-madamanchi-80818b5/
Undergraduate Student Alumni
